ChimeraX CircosContacts
A ChimeraX plugin that converts predicted contact sets into interactive circos-style maps for analysis and visualization.
A ChimeraX plugin that converts predicted contact sets into interactive circos-style maps for analysis and visualization.
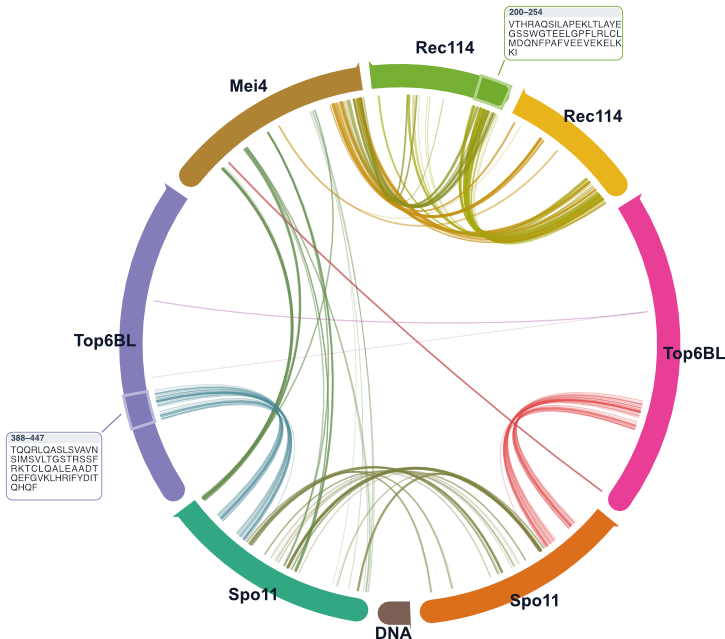
Detailed user docs are available for command semantics, interactive web controls, workflows, and troubleshooting.
circoscontacts on open or selected models.contacts-style selection and restrict semantics.Choose open models (or subsets) using standard ChimeraX atom spec syntax.
Plugin runs ChimeraX contacts with your source/restrict settings.
Adjust chain order, orientation, thresholds, callouts, and region annotations.
Save SVG, ChimeraX color script (.cxc), and session (.json).
Peter Carlton, www.carltonlab.org